Dataverse
Main Page > Documentation > Requirements > Dataverse
This page tracks development of a proof of concept integration of Archivematica with Dataverse. Please note that this feature is experimental and Artefactual has not tested it in a production environment.
See also
Overview
This wiki captures requirements for ingesting studies (datasets) from Dataverse into Archivematica for long-term preservation.
Installation
April 24, 2017 This feature requires a development branch of Archivematica, which can be installed with the following steps:
1) install deploy-pub. https://github.com/artefactual/deploy-pub 2) use the archivematica-centos7 playbook in deploy-pub https://github.com/artefactual/deploy-pub/tree/master/playbooks/archivematica-centos7 3) create a hosts file that lists your target machine (see digital ocean example linked from playbook) 4) in requirements.yml change version of ansible-archivematica-src to "stable/1.6.x" 5) change singlenode.yml to point to the host you defined in your hosts file. 6) change the vars-singlenode.yml to include the following info:
- required for dataverse testing
archivematica_src_am_version: "dev/dataverse-poc" archivematica_src_automationtools: "yes" archivematica_src_automationtools_version: "dev/dataverse"
Workflow
- The proposed workflow consists of issuing API calls to Dataverse, receiving content (data files and metadata) for ingest into Archivematica, preparing Archivematica Archival Information Packages (AIPs) and placing them in archival storage,
and updating the Dataverse study with the AIP UUIDs(this was determined to be out of scope). - Analysis is based on Dataverse tests using https://apitest.dataverse.org and https://demo.dataverse.org, online documentation at http://guides.dataverse.org/en/latest/api/index.html and discussions with Dataverse developers and users.
- Proposed integration is for Archivematica 1.5 and higher and Dataverse 4.x.
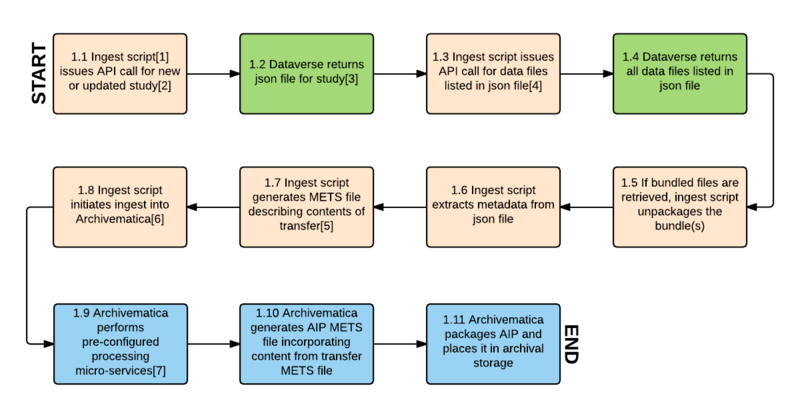
Workflow diagram
Workflow diagram notes
[1] "Ingest script" refers to an automation tool designed to automate ingest into Archivematica for bulk processing. An existing automation tool would be modified to accomplish the tasks described in the workflow.
[2] A new or updated study is one that has been published, either for the first time or as a new version, since the last API call.
[3] The json file contains citation and other study-level metadata, an entity_id field that is used to identify the study in Dataverse, version information, a list of data files with their own entity_id values, and md5 checksums for each data file.
[4] If json file has content_type of tab separated values, Archivematica issues API call for multiple file ("bundled") content download. This returns a zipped package for tsv files containing the .tab file, the original uploaded file, several other derivative formats, a DDI XML file and file citations in Endnote and RIS formats.
[5] The METS file will consist of a dmdSec containing the DC elements extracted from the json file, and a fileSec and structMap indicating the relationships between the files in the transfer (eg. original uploaded data file, derivative files generated for tabular data, metadata/citation files). This will allow Archivematica to apply appropriate preservation micro-services to different filetypes and provide an accurate representation of the study in the AIP METS file (step 1.9).
[6] Archivematica ingests all content returned from Dataverse, including the json file, plus the METS file generated in step 1.6.
[7] Standard and pre-configured micro-services include: assign UUID, verify checksums, generate checksums, extract packages, scan for viruses, clean up filenames, identify formats, validate formats, extract metadata and normalize for preservation.
Transfer METS file
When the ingest script retrieves content from Dataverse, it generates a METS file to allow Archivematica to understand the contents of the transfer and the relationships between its various data and metadata files.
Sample transfer METS file
Original Dataverse study retrieved through API call:
- dataset.json (a JSON file generated by Dataverse consisting of study-level metadata and information about data files)
- Study_info.pdf (a non-tabular data file)
- A zipped bundle consisting of the following:
- YVR_weather_data.sav (an SPSS SAV file uploaded by the researcher)
- YVR_weather_data.tab (a TAB file generated from the SPSS SAV file by Dataverse)
- YVR weather_data.RData (an R file generated from the SPSS SAV file by Dataverse)
- YVR_weather_data-ddi.xml, YVR_weather_datacitation-endnote.xml, and YVR_weather_datacitation-ris.ris (three metadata files generated for the TAB file by Dataverse)
Resulting transfer METS file
- The fileSec in the METS file consists of three file groups, USE="original" (the PDF and SAV files); USE="derivative" (the TAB and R files); and USE="metadata" (the JSON file and the three metadata files from the zipped bundle).
- All of the files unpacked from the Dataverse bundle have a GROUPID attribute to indicate the relationship between them. If the transfer had consisted of more than one bundle, each set of unpacked files would have its own GROUPID.
- Three dmdSecs have been generated:
- dmdSec_1, consisting of a small number of study-level DDI terms
- dmdSec_2, consisting of an mdRef to the JSON file
- dmdSec_3, consisting of an mdRef to the DDI XML file
- In the structMap, dmdSec_1 and dmdSec_2 are linked to the study as a whole, while dmdSec_3 is linked to the TAB file. The endnote and ris files have not been made into dmdSecs because they contain small subsets of metadata which are already captured in dmdSec_1 and the DDI xml file.
Metadata sources for METS file
| METS element | Information source | Notes |
|---|---|---|
| ddi:titl | json: citation/typeName: "title", value: [value] | |
| ddi:IDNo | json: authority, identifier | json example: "authority": "10.5072/FK2/", "identifier": "0MOPJM" |
| ddi:IDNo agency attribute | json: protocol | json example: "protocol": "doi" |
| ddi:AuthEntity | json: citation/typeName: "authorName" | |
| ddi:distrbtr | Config setting in ingest tool | |
| ddi:version date attribute | json: "releaseTime" | |
| ddi:version type attribute | json: "versionState" | |
| ddi:version | json: "versionNumber", "versionMinorNumber" | |
| ddi:restrctn | json: "termsOfUse" | |
| fileGrp USE="original" | json: datafile | Each non-tabular data file is listed as a datafile in the files section. Each TAB file derived by Dataverse for uploaded tabular file formats is also listed as a datafile, with the original file uploaded by the researcher indicated by "originalFileFormat". |
| fileGrp USE="derivative" | All files that are included in a bundle, except for the original file and the metadata files (see below). | |
| fileGrp USE="metadata" | Any files with .json or .ris extension, any -ddi.xml files and -endnote.xml files | |
| CHECKSUM | json: datafile/"md5": [value] | |
| CHECKSUMTYPE | json: datafile/"md5" | |
| GROUPID | Generated by ingest tool. Each file unpacked from a bundle is given the same group id. |
AIP METS file
Basic METS file structure
The Archival Information Package (AIP) METS file will follow the basic structure for a standard Archivematica AIP METS file described at METS. A new fileGrp USE="derivative" will be added to indicate TAB, RData and other derivatives generated by Dataverse for uploaded tabular data format files.
dmdSecs in AIP METS file
The dmdSecs in the transfer METS file will be copied over to the AIP METS file.
Additions to PREMIS for derivative files
In the PREMIS Object entity, relationships between original and derivative tabular format files from Dataverse will be described using PREMIS relationship semantic units. A PREMIS derivation event will be added to indicate the derivative file was generated from the original file, and a Dataverse Agent will be added to indicate the Event were carried out by Dataverse prior to ingest, rather than by Archivematica.
Note We originally considered adding a creation event for the derivative files as well, but decided that it's not necessary as the event can be inferred from the derivation event and the PREMIS object relationships.
Note "Derivation" is not an event type on the Library of Congress controlled vocabulary list at http://id.loc.gov/vocabulary/preservation/eventType.html. However, we have submitted it as a proposed new term at http://premisimplementers.pbworks.com/w/page/102413902/Preservation%20Events%20Controlled%20Vocabulary - a list of new terms that is being considered by the PREMIS Editorial Committee.
Example:
Original SPSS SAV file
<premis:relationship>
<premis:relationshipType>derivation</premis:relationshipType>
<premis:relationshipSubType>is source of</premis:relationshipSubType>
<premis:relatedObjectIdentification>
<premis:relatedObjectIdentifierType>UUID</premis:relatedObjectIdentifierType>
<premis:relatedObjectIdentifierValue>[TAB file UUID]</premis:relatedObjectIdentifierValue>
<premis:relationship>
...
<premis:eventIdentifier>
<premis:eventIdentifierType>UUID</premis:eventIdentifierType>
<premis:eventIdentifierValue>[Event UUID assigned by Archivematica]</premis:eventIdentifierValue>
</premis:eventIdentifier>
<premis:eventType>derivation</premis:eventType>
<premis:eventDateTime>2015-08-21</premis:eventDateTime>
<premis:linkingAgentIdentifier>
<premis:linkingAgentIdentifierType>URI</premis:linkingAgentIdentifierType>
<premis:linkingAgentIdentifierValue>http://dataverse.scholarsportal.info/dvn/
</premis:linkingAgentIdentifierValue>
</premis:linkingAgentIdentifier>
...
<premis:agentIdentifier>
<premis:agentIdentifierType>URI</premis:agentIdentifierType>
<premis:agentIdentifierValue>http://dataverse.scholarsportal.info/dvn/</premis:agentIdentifierValue>
</premis:agentIdentifier>
<premis:agentName>SP Dataverse Network</premis:agentName>
<premis:agentType>organization</premis:agentType>
Derivative TAB file
<premis:relationship>
<premis:relationshipType>derivation</premis:relationshipType>
<premis:relationshipSubType>has source</premis:relationshipSubType>
<premis:relatedObjectIdentification>
<premis:relatedObjectIdentifierType>UUID</premis:relatedObjectIdentifierType>
<premis:relatedObjectIdentifierValue>[SPSS SAV file UUID]</premis:relatedObjectIdentifierValue>
<premis:relationship>
Fixity check for checksums received from Dataverse
<premis:eventIdentifier>
<premis:eventIdentifierType>UUID</premis:eventIdentifierType>
<premis:eventIdentifierValue>[Event UUID assigned by Archivematica]</premis:eventIdentifierValue>
</premis:eventIdentifier>
<premis:eventType>fixity check</premis:eventType>
<premis:eventDateTime>2015-08-21</premis:eventDateTime>
<premis:eventDetail>program="python"; module="hashlib.sha256()"</premis:eventDetail>
<premis:eventOutcomeInformation>
<premis:eventOutcome>Pass</premis:EventOutcome>
<premis:eventOutcomeDetail>
<premis:eventOutcomeDetailNote>Dataverse checksum 91b65277959ec273763d28ef002e83a6b3fba57c7a3[...]
verified</premis:eventOutcomeDetailNote>
</premis:eventOutcomeDetail>
<premis:eventOutcomeInformation>
</premis:linkingAgentIdentifier>
<premis:linkingAgentIdentifierType>preservation system</premis:linkingAgentIdentifierType>
<premis:linkingAgentIdentifierValue>Archivematica 1.4.1</premis:linkingAgentIdentifierValue>
</premis:linkingAgentIdentifier>
AIP structure
An Archival Information Package derived from a Dataverse ingest will have the same basic structure as a generic Archivematica AIP, described at AIP_structure. There are additional metadata files that are included in a Dataverse-derived AIP, and each zipped bundle that is included in the ingest will result in a separate directory in the AIP. The following is a sample structure.
Bag structure
The Archival Information Package (AIP) is packaged in the Library of Congress BagIt format, and may be stored compressed or uncompressed:
Pacific_weather_patterns_study-dfb0b75d-6555-4e99-a8d8-95bed0f6303f.7z ├── bag-info.txt ├── bagit.txt ├── manifest-sha512.txt│ ├── tagmanifest-md5.txt └── data [standard bag directory containing contents of the AIP]
AIP structure
All of the contents of the AIP reside within the data directory:
├── data │ ├── logs [log files generated during processing] │ │ ├── fileFormatIdentification.log │ │ └── transfers │ │ └── Pacific_weather_patterns_study-1a0f309a-d3ec-43ee-bb48-a868cd5ca85c │ │ └── logs │ │ ├── extractContents.log │ │ ├── fileFormatIdentification.log │ │ └── filenameCleanup.log │ ├── METS.dfb0b75d-6555-4e99-a8d8-95bed0f6303f.xml [the AIP METS file] │ ├── objects [a directory containing the digital objects being preserved, plus their metadata] │ ├── chelan_052.jpg [an original file from Dataverse] │ ├── Weather_data.sav [an original file from Dataverse] │ ├── Weather_data [a bundle retrieved from Dataverse] │ │ ├── Weather_data.xml │ │ ├── Weather_data.ris │ │ ├── Weather_data-ddi.xml │ │ └── Weather_data.tab [a TAB derivative file generated by Dataverse] │ ├── metadata │ │ └── transfers │ │ └── Pacific_weather_patterns_study-1a0f309a-d3ec-43ee-bb48-a868cd5ca85c │ │ ├── agents.json [information about the source of the data, used to populate the PREMIS Dataverse agent in the AIP METS file] │ │ ├── dataset.json [the full json file retrieved from Dataverse] │ │ └── METS.xml [the METS file generated by the ingest script to prepare Dataverse contents for ingest into Archivematica] │ └── submissionDocumentation │ └── transfer-58-1a0f309a-d3ec-43ee-bb48-a868cd5ca85c │ └── METS.xml [a standard transfer METS file generated to list all contents of an Archivematica transfer]
AIP METS file structure
The AIP METS file records information a bout the contents of the AIP, and indicates the relationships between the various files in the AIP. A sample AIP METS file would be structured as follows:
METS header -Date METS file was created METS dmdSec [descriptive metadata section] -DDI XML metadata taken from the METS transfer file, as follows --ddi:title --ddi:IDno --ddi:authEnty --ddi:distrbtr --ddi:version --ddi:restrctn METS dmdSec [descriptive metadata section] -link to dataset.json METS dmdSec [descriptive metadata section] -link to DDI.XML file created for derivative file as part of bundle METS amdSec [administrative metadata section, one for each original, derivative and normalized file in the AIP] -techMD [technical metadata] --PREMIS technical metadata about a digital object, including file format information and extracted metadata -digiprovMD [digital provenance metadata] --PREMIS event: derivation (for derived formats) -digiprovMD [digital provenance metadata] --PREMIS event:ingestion -digiprovMD [digital provenance metadata] --PREMIS event: unpacking (for bundled files) -digiprovMD [digital provenance metadata] --PREMIS event: message digest calculation -digiprovMD [digital provenance metadata] --PREMIS event: virus check -digiprovMD [digital provenance metadata] --PREMIS event: format identification -digiprovMD [digital provenance metadata] --PREMIS event: fixity check (if file comes from Dataverse with a checksum) -digiprovMD [digital provenance metadata] --PREMIS event: normalization (if file is normalized to a preservation format during Archivematica processing) -digiprovMD [digital provenance metadata] --PREMIS event: creation (if file is a normalized preservation master generated during Archivematica processing) -digiprovMD --PREMIS agent: organization -digiprovMD --PREMIS agent: software -digiprovMD --PREMIS agent: Archivematica user METS fileSec [file section] -fileGrp USE="original" [file group] --original files uploaded to Dataverse -fileGrp USE="derivative" --derivative tabular files generated by Dataverse -fileGrp USE="submissionDocumentation" --METS.XML (standard Archivematica transfer METS file listing contents of transfer) -fileGrp USE="preservation" --normalized preservation masters generated during Archivematica processing -fileGrp USE="metadata" --dataset.json --DDI.XML --xcitation-endnote.xml --xcitation-ris.ris METS structMap [structural map] -directory structure of the contents of the AIP